Phytoplankton diversity in a tropical bay, North Borneo, Malaysia as revealed by light microscopy and Next-Generation Sequencing
-
Abstract: Assessments of phytoplankton diversity in Sabah waters, North Borneo, have primarily relied on morphology-based identification, which has inherent biases and can be time-consuming. Next-Generation Sequencing (NGS) technology has been shown to be capable of overcoming several limitations of morphology-based methods. Samples were collected from the Sepanggar Bay over the course of the year 2018 in different monsoon seasons. Morphology-based identification and NGS sequencing of the V8–V9 region of the 18S LSU rDNA were used to investigate the diversity of the phytoplankton community. Microscopy and NGS showed complementary results with more diatom taxa detected by microscopy whereas NGS detected smaller and rarer taxa. The harmful algal genera in the study site comprised of Skeletonema, Margalefidinium, Pyrodinium, Takayama, and Alexandrium as detected by NGS. This study showed that that an integrative approach of both morphological and molecular techniques could provide more comprehensive information about the phytoplankton community as the approach captured quantitative variability as well as the diversity of phytoplankton species.
-
Key words:
- phytoplankton community /
- Next-Generation Sequencing (NGS) /
- Sabah /
- South China Sea
-
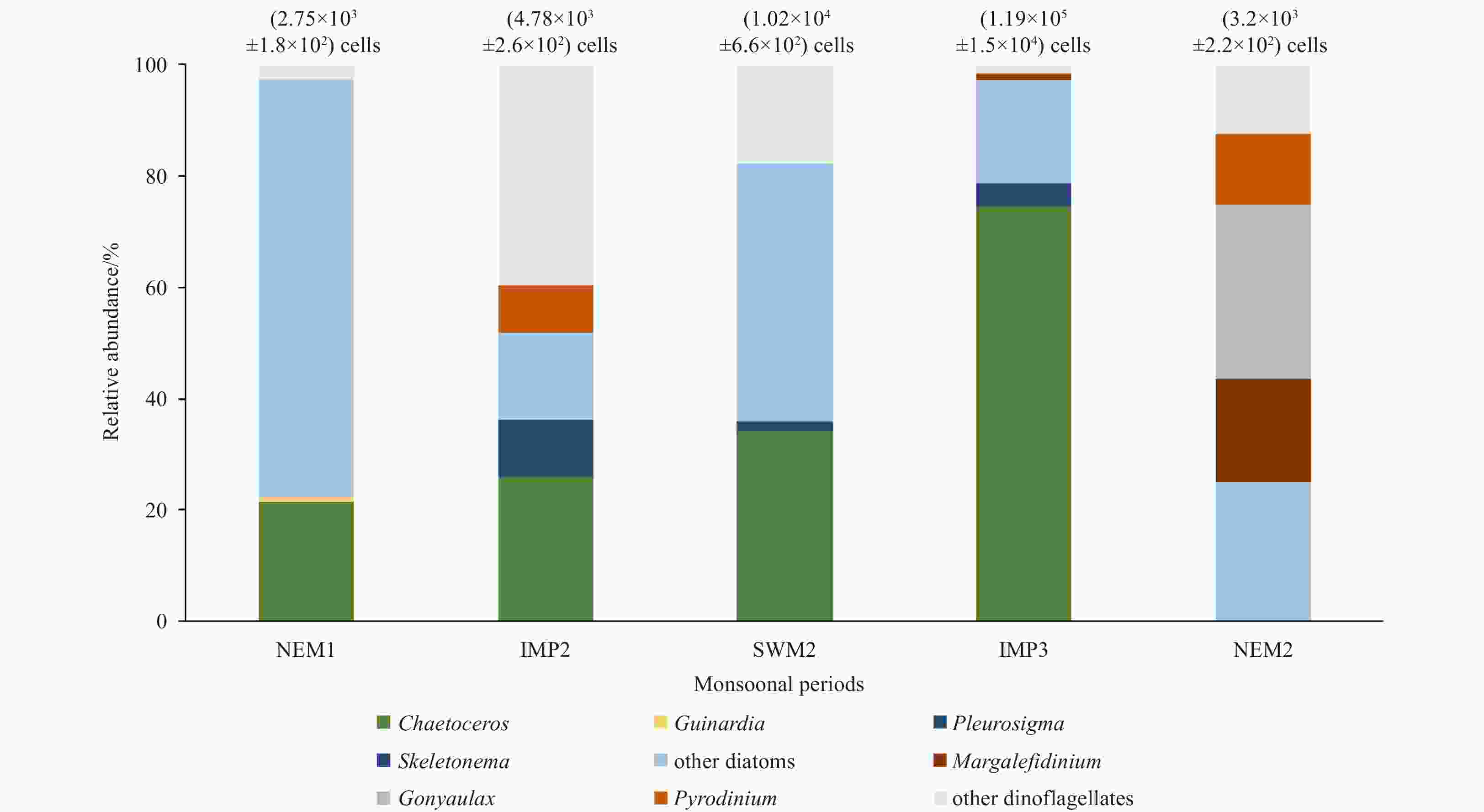
Figure 3. Bar chart of relative abundances of phytoplankton based on cell counts. Total cell counts are provided in brackets above the respective bars. The group “other diatoms” include Asteromphalus, Bacteriastrum, Coscinodiscus, Cylindrotheca, Dactyliosolen, Ditylum, Eucampia, Lauderia, Leptocylindrus, Navicula, Nitzschia, Rhizosolenia, Thalassionema, and Pseudo-nitzschia. The group “other dinoflagellates” include Akashiwo, Ceratium, Dinophysis, Gymnodinium, Gyrodinium, and Prorocentrum.
Table 1. Taxonomic composition as revealed by amplicon sequencing
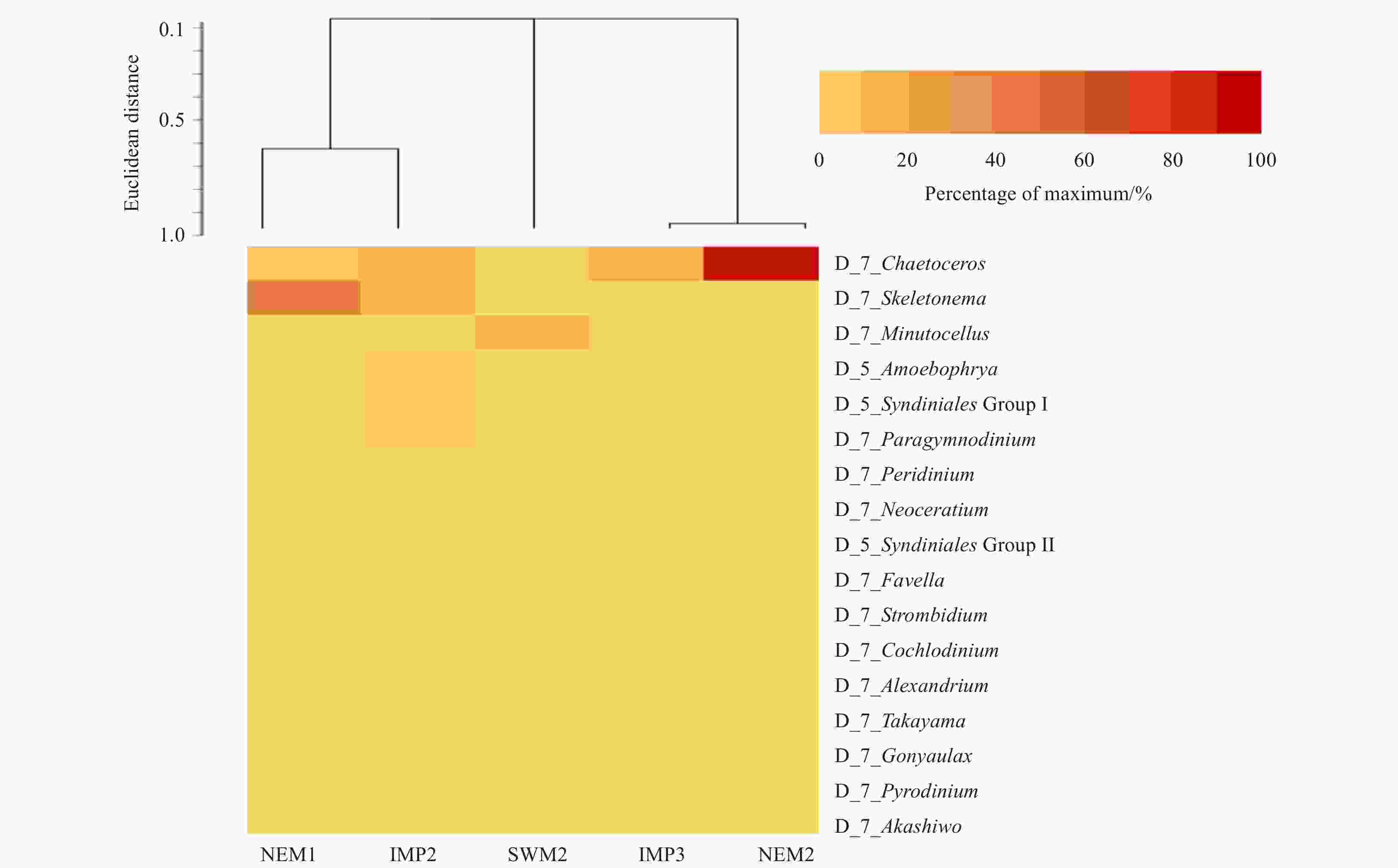
Group Rank Taxon Relative abundance/% NEM1 IMP2 SWM2 IMP3 NEM2 Diatom Subphylum Coscinodiscophytina 0.0 0.0 0.0 0.0 <0.1 Class Bacillariophyceae 0.9 0.0 0.0 4.3 1.0 Class Mediophyceae <0.1 <0.1 <0.1 1.4 0.5 Order Fragilariales 0.0 0.0 0.0 <0.1 <0.1 Genus Chaetoceros 17.3 22.0 2.5 55.7 87.1 Genus Corethron <0.1 0.0 0.0 <0.1 <0.1 Genus Coscinodiscus 0.0 0.0 0.0 <0.1 <0.1 Genus Cyclotella 0.9 1.0 2.5 1.4 1.0 Genus Leptocylindrus <0.1 <0.1 <0.1 1.4 <0.1 Genus Minutocellus 1.8 3.0 72.5 <0.1 1.0 Genus Navicula <0.1 0.0 0.0 <0.1 <0.1 Genus Odontella 0.0 0.0 0.0 <0.1 <0.1 Genus Rhizosolenia <0.1 <0.1 0.0 <0.1 0.0 Genus Skeletonema 63.6 14.0 2.5 7.1 1.9 Genus Thalassiosira <0.1 1.0 0.0 <0.1 <0.1 Dinoflagellate Infraphylum Dinoflagellata 2.7 12.0 5.0 4.3 1.0 Sub-class Gymnodiniphycidae <0.1 <0.1 <0.1 <0.1 <0.1 Order Dinophysiales 0.0 0.0 <0.1 <0.1 <0.1 Order Gonyaulacales 0.9 0.0 0.0 <0.1 0.0 Genus Peridiniales 0.0 0.0 0.0 <0.1 <0.1 Family Suessiaceae <0.1 <0.1 0.0 0.0 <0.1 Family Thoracosphaeraceae <0.1 0.0 0.0 <0.1 0.0 Genus Akashiwo 0.0 <0.1 0.0 0.0 0.0 Genus Alexandrium <0.1 <0.1 <0.1 1.4 <0.1 Genus Amphidiniopsis 0.0 0.0 <0.1 0.0 <0.1 Genus Amphidinium 0.9 <0.1 <0.1 <0.1 <0.1 Genus Azadinium 0.0 0.0 <0.1 0.0 0.0 Genus Blixaea <0.1 <0.1 <0.1 8.6 1.9 Genus Cochlodinium <0.1 <0.1 <0.1 2.9 0.5 Genus Diplosalis 0.0 0.0 0.0 <0.1 0.0 Genus Erythropsidinium 0.0 0.0 <0.1 2.9 <0.1 Genus Fragilidium <0.1 0.0 0.0 <0.1 <0.1 Genus Gonyaulax <0.1 1.0 <0.1 <0.1 <0.1 Genus Gymnodinium 0.9 <0.1 <0.1 <0.1 0.5 Genus Gyrodinium 1.8 0.0 <0.1 2.9 <0.1 Genus Haplozoon <0.1 1.0 <0.1 <0.1 <0.1 Genus (cyst) Islandinium 0.0 0.0 0.0 <0.1 0.0 Genus Katodinium 0.0 <0.1 0.0 <0.1 <0.1 Genus Oodinium <0.1 <0.1 5.0 0.0 <0.1 Genus Paragymnodinium 0.9 6.0 2.5 <0.1 <0.1 Genus Polykrikos 0.0 <0.1 0.0 <0.1 <0.1 Genus Posoniella 0.0 <0.1 0.0 0.0 0.0 Genus Prorocentrum 0.0 0.0 0.0 <0.1 <0.1 Genus Protoperidinium <0.1 1.0 <0.1 1.4 0.5 Genus Ptychodiscus <0.1 <0.1 <0.1 <0.1 <0.1 Genus Pyrodinium <0.1 0.0 <0.1 <0.1 0.5 Genus Sinophysis <0.1 0.0 <0.1 <0.1 <0.1 Genus Symbiodinium 0.9 1.0 <0.1 <0.1 0.5 Genus Takayama <0.1 1.0 <0.1 <0.1 <0.1 Genus Tripos 4.5 1.0 <0.1 1.4 0.5 Genus Woloszynskia <0.1 0.0 0.0 <0.1 <0.1 Clade Gymnodinium clade 1.8 <0.1 2.5 2.8 1.4 Eustigmatophyte Order Eustigmatales <0.1 <0.1 <0.1 <0.1 0.0 Ochrophyte Phylum Ochrophyta 0.9 5.0 2.5 1.4 0.5 Raphidophyte Order Chattonellales <0.1 <0.1 <0.1 <0.1 <0.1 Class Raphidophyceae <0.1 <0.1 <0.1 <0.1 <0.1 Silicoflagellate Class Dictyophyceae 0.0 <0.1 0.0 <0.1 <0.1 Stramenopile Genus Phaeomonas 0.0 1.0 0.0 0.0 0.0 Genus Pinguiochrysis 0.0 <0.1 0.0 0.0 0.0 Chrysophyte Class Chrysophyceae 0.0 <0.1 <0.1 <0.1 <0.1 Genus Ankylochrisis 0.0 0.0 0.0 0.0 <0.1 Genus Chrysowaernella 0.0 0.0 0.0 <0.1 0.0 Genus Ochromonas 0.0 0.0 0.0 0.0 <0.1 Genus Paraphysomonas 0.0 0.0 0.0 <0.1 <0.1 Cryptophyte Class Cryptophyceae <0.1 0.0 <0.1 <0.1 <0.1 Order Cryptomonadales <0.1 2.0 <0.1 <0.1 0.5 Genus Leucocryptos <0.1 1.0 <0.1 <0.1 <0.1 Genus Proteomonas 0.9 1.0 <0.1 <0.1 <0.1 Genus Teleaulax 0.0 <0.1 <0.1 0.0 <0.1 Species Leucocryptos marina <0.1 <0.1 0.0 <0.1 <0.1 Species Hemiselmis virescens 0.0 0.0 0.0 0.0 <0.1 Species Proteomonas sulcata <0.1 1.0 <0.1 <0.1 <0.1 Chlorophyte Phylum Chlorophyta 0.9 7.0 5.0 <0.1 1.0 Class Mamiellophyceae <0.1 <0.1 <0.1 0.0 <0.1 Order Chlorodendrales <0.1 2.0 7.5 1.4 <0.1 Order Mamiellales <0.1 8.0 <0.1 <0.1 <0.1 Order Ulvales <0.1 <0.1 <0.1 <0.1 <0.1 Genus Bathycoccus 0.0 0.0 0.0 <0.1 0.0 Genus Coccomyxa 0.0 <0.1 0.0 <0.1 <0.1 Genus Crustomastix 0.0 <0.1 0.0 0.0 0.0 Genus Mamiella <0.1 <0.1 <0.1 0.0 <0.1 Genus Nephroselmis <0.1 2.0 <0.1 <0.1 <0.1 Genus Prasinoderma 0.0 <0.1 0.0 0.0 <0.1 Genus Tetraselmis 0.0 <0.1 <0.1 0.0 <0.1 Total phytoplankton reads 10 046 8 363 3 869 6 331 18 745 Total reads 88 124 88 028 87 927 87 928 88 007 Unassigned reads 1 374 20 327 128 189 1 026 Note: Values with “<” indicate sequences that have at least one read. -
Ajani P A, Verma A, Kim J H, et al. 2021. Using qPCR and high-resolution sensor data to model a multi-species Pseudo-nitzschia (Bacillariophyceae) bloom in southeastern Australia. Harmful Algae, 108: 102095. doi: 10.1016/j.hal.2021.102095 Anton A, Teoh P L, Mohd-Shaleh S R, et al. 2008. First occurrence of Cochlodinium blooms in Sabah, Malaysia. Harmful Algae, 7(3): 331–336. doi: 10.1016/j.hal.2007.12.013 Band-Schmidt C J, Martínez-López A, Bustillos-Guzmán J J, et al. 2012. Morphology, biochemistry, and growth of raphidophyte strains from the Gulf of California. Hydrobiologia, 693(1): 81–97. doi: 10.1007/s10750-012-1088-y Bates S S, Hubbard K A, Lundholm N, et al. 2018. Pseudo-nitzschia, Nitzschia, and domoic acid: new research since 2011. Harmful Algae, 79: 3–43. doi: 10.1016/j.hal.2018.06.001 Bazin P, Jouenne F, Friedl T, et al. 2014. Phytoplankton diversity and community composition along the estuarine gradient of a temperate macrotidal ecosystem: combined morphological and molecular approaches. PLoS ONE, 9(4): e94110. doi: 10.1371/journal.pone.0094110 Bergman B, Sandh G, Lin Senjie, et al. 2013. Trichodesmium– a widespread marine cyanobacterium with unusual nitrogen fixation properties. FEMS Microbiology Reviews, 37(3): 286–302. doi: 10.1111/j.1574-6976.2012.00352.x Boenigk J, Pfandl K, Stadler P, et al. 2005. High diversity of the ‘Spumella-like’ flagellates: an investigation based on the SSU rRNA gene sequences of isolates from habitats located in six different geographic regions. Environmental Microbiology, 7(5): 685–697. doi: 10.1111/j.1462-2920.2005.00743.x Bowers H A, Tengs T, Glasgow H B, et al. 2000. Development of real-time PCR assays for rapid detection of Pfiesteria piscicida and related dinoflagellates. Applied and Environmental Microbiology, 66(11): 4641–4648. doi: 10.1128/AEM.66.11.4641-4648.2000 Bradley I M, Pinto A J, Guest J S. 2016. Design and evaluation of illumina MiSeq-compatible, 18S rRNA gene-specific primers for improved characterization of mixed phototrophic communities. Applied and Environmental Microbiology, 82(19): 5878–5891. doi: 10.1128/AEM.01630-16 Carpenter E J, Montoya J P, Burns J, et al. 1999. Extensive bloom of a N2-fixing diatom/cyanobacterial association in the tropical Atlantic Ocean. Marine Ecology Progress Series, 185: 273–283. doi: 10.3354/meps185273 Chen Zhenfan, Zhang Qingchun, Kong Fanzhou, et al. 2019. Resolving phytoplankton taxa based on high-throughput sequencing during brown tides in the Bohai Sea, China. Harmful Algae, 84: 127–138. doi: 10.1016/j.hal.2019.03.011 Chong B W K, Leong S C Y, Kuwahara V S, et al. 2020. Monsoonal variation of the marine phytoplankton community in Kota Kinabalu, Sabah. Regional Studies in Marine Science, 37: 101326. doi: 10.1016/j.rsma.2020.101326 Cupp E E. 1943. Marine plankton diatoms of the west coast of north America. Bulletin of the Scripps Institution of Oceanography, 5(1): 199–207 de Salas M F, Bolch C J S, Botes L, et al. 2003. Takayama gen. nov. (Gymnodiniales, Dinophyceae), a new genus of unarmored dinoflagellates with sigmoid apical grooves, including the description of two new species. Journal of Phycology, 39(6): 1233–1246. doi: 10.1111/j.0022-3646.2003.03-019.x de Salas M F, Rhodes L L, Mackenzie L A, et al. 2005. Gymnodinoid genera Karenia and Takayama (Dinophyceae) in New Zealand coastal waters. New Zealand Journal of Marine and Freshwater Research, 39(1): 135–139. doi: 10.1080/00288330.2005.9517296 Edvardsen B, Eikrem W, Green J C, et al. 2000. Phylogenetic reconstructions of the Haptophyta inferred from 18S ribosomal DNA sequences and available morphological data. Phycologia, 39(1): 19–35. doi: 10.2216/i0031-8884-39-1-19.1 Elferink S, Neuhaus S, Wohlrab S, et al. 2017. Molecular diversity patterns among various phytoplankton size-fractions in west Greenland in late summer. Deep-Sea Research Part I: Oceanographic Research Papers, 121: 54–69. doi: 10.1016/j.dsr.2016.11.002 Furuya K, Iwataki M, Lim P T, et al. 2018. Overview of harmful algal blooms in Asia. In: Glibert P M, Berdalet E, Burford M A, et al., eds. Global Ecology and Oceanography of Harmful Algal Blooms. Cham: Springer, 289–308 Galluzzi L, Penna A, Bertozzini E, et al. 2004. Development of a real-time PCR assay for rapid detection and quantification of Alexandrium minutum (a dinoflagellate). Applied and Environmental Microbiology, 70(2): 1199–1206. doi: 10.1128/AEM.70.2.1199-1206.2004 Giribet G, Wheeler W. 2005. On bivalve phylogeny: a high-level analysis of the Bivalvia (Mollusca) based on combined morphology and DNA sequence data. Invertebrate Biology, 121(4): 271–324. doi: 10.1111/j.1744-7410.2002.tb00132.x Gómez F, Richlen M L, Anderson D M. 2017. Molecular characterization and morphology of Cochlodinium strangulatum, the type species of Cochlodinium, and Margalefidinium gen. nov. for C. polykrikoides and allied species (Gymnodiniales, Dinophyceae). Harmful Algae, 63: 32–44. doi: 10.1016/j.hal.2017.01.008 Gran-Stadniczeñko S, Egge E, Hostyeva V, et al. 2019. Protist diversity and seasonal dynamics in Skagerrak plankton communities as revealed by metabarcoding and microscopy. Journal of Eukaryotic Microbiology, 66(3): 494–513. doi: 10.1111/jeu.12700 Hartley B. 1996. An Atlas of British Diatoms. Avon Dassett: Biopress Hii K S, Law I K, Sing L W L, et al. 2019. Wide distribution of toxic marine dinoflagellate Alexandrium tamiyavanichii along the east coast of Peninsular Malaysia. In: Akhir M F M, ed. National Scientific Cruise Expedition 2016–2017. Lahore: UMT Press, 101–113 Hii K S, Mohd-Din M, Luo Zhaohe, et al. 2021. Diverse harmful microalgal community assemblages in the Johor Strait and the environmental effects on its community dynamics. Harmful Algae, 107: 102077. doi: 10.1016/j.hal.2021.102077 Hong D D, Hien H T M, Thu N H, et al. 2008. Phylogenetic analyses of Prorocentrum spp. and Alexandrium spp. isolated from northern coast of Vietnam based on 18S rDNA sequence. Journal of Environmental Biology, 29(4): 535–542 Hosoi-Tanabe S, Sako Y. 2005. Species-specific detection and quantification of toxic marine dinoflagellates Alexandrium tamarense and A. catenella by real-time PCR assay. Marine Biotechnology, 7(5): 506–514. doi: 10.1007/s10126-004-4128-4 Iwataki M, Kawami H, Mizushima K, et al. 2008. Phylogenetic relationships in the harmful dinoflagellate Cochlodinium polykrikoides (Gymnodiniales, Dinophyceae) inferred from LSU rDNA sequences. Harmful Algae, 7(3): 271–277. doi: 10.1016/j.hal.2007.12.003 Jipanin S J, Shaleh S R M, Lim P T, et al. 2019. The monitoring of harmful algae blooms in Sabah, Malaysia. Journal of Physics: Conference Series, 1358(1): 012014. doi: 10.1088/1742-6596/1358/1/012014 Kadar N A, Raehanah S, Shaleh M, et al. 2018. Molecular and phylogenetic identification of marine microalgae inferred by 18S rDNA gene. Malaysian Applied Biology, 47(6): 41–45 Kon N F, Lau W L S, Hii K S, et al. 2017. Quantitative real-time PCR detection of a harmful unarmoured dinoflagellate, Karlodinium australe (Dinophyceae). Phycological Research, 65(4): 291–298. doi: 10.1111/pre.12186 Kon N F, Teng S T, Hii K S, et al. 2015. Spatial distribution of toxic Alexandrium tamiyavanichii (Dinophyceae) in the southeastern South China Sea-Sulu Sea: a molecular-based assessment using real-time quantitative PCR (qPCR) assay. Harmful Algae, 50: 8–20. doi: 10.1016/j.hal.2015.10.002 Leong S, Lim L P, Chew S M, et al. 2015. Three new records of dinoflagellates in Singapore’s coastal waters, with observations on environmental conditions associated with microalgal growth in the Johor Straits. The Raffles Bulletin of Zoology, S31: 24–36 Lewandowska A M, Breithaupt P, Hillebrand H, et al. 2012. Responses of primary productivity to increased temperature and phytoplankton diversity. Journal of Sea Research, 72: 87–93. doi: 10.1016/j.seares.2011.10.003 Lim H C, Leaw C P, Tan T H, et al. 2014. A bloom of Karlodinium australe (Gymnodiniales, Dinophyceae) associated with mass mortality of cage-cultured fishes in west Johor Strait, Malaysia. Harmful Algae, 40: 51–62. doi: 10.1016/j.hal.2014.10.005 Lim Zhenfei, Luo Zhaohe, Lee L K, et al. 2019. Taxonomy and toxicity of Prorocentrum from Perhentian Islands (Malaysia), with a description of a non-toxigenic species Prorocentrum malayense sp. nov. (Dinophyceae). Harmful Algae, 83: 95–108. doi: 10.1016/j.hal.2019.01.007 Lim P T, Usup G, Leaw C P. 2012. Harmful algal blooms in Malaysian waters. Sains Malaysiana, 41(12): 1509–1515 Liow G R, Lau W L S, Law I K, et al. 2019. Phytoplankton community changes in Kuantan Port (Malaysia), with emphasis on the paralytic-shellfish toxin-producing dinoflagellate Alexandrium tamiyavanichii. Regional Studies in Marine Science, 26: 100504. doi: 10.1016/j.rsma.2019.100504 Liu Hui, Probert I, Uitz J, et al. 2009. Extreme diversity in noncalcifying haptophytes explains a major pigment paradox in open oceans. Proceedings of the National Academy of Sciences of the United States of America, 106(31): 12803–12808. doi: 10.1073/pnas.0905841106 Lohan K M P, Fleischer R C, Carney K J, et al. 2016. Amplicon-based pyrosequencing reveals high diversity of protistan parasites in ships’ ballast water: implications for biogeography and infectious diseases. Microbial Ecology, 71(3): 530–542. doi: 10.1007/s00248-015-0684-6 López-García P, Rodríguez-Valera F, Pedrós-Alió C, et al. 2001. Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature, 409(6820): 603–607. doi: 10.1038/35054537 Lum W M, Benico G, Azanza R, et al. 2019. Morphology and molecular phylogeny of the harmful raphidophyte Chattonella subsalsa isolated from Bolinao, Philippines. Philippine Journal of Natural Sciences, 24: 50–56 Lund J W, Hendey N I. 1965. An introductory account of the smaller algae of British coastal waters. Part V. Bacillariophyceae (Diatoms). Journal of Ecology, 53(2): 549 Menden-Deuer S, Lessard E J, Satterberg J. 2001. Effect of preservation on dinoflagellate and diatom cell volume and consequences for carbon biomass predictions. Marine Ecology Progress Series, 222: 41–50. doi: 10.3354/meps222041 Mohammad-Noor N, Weliyadi E, Aung T, et al. 2014. Effects of meteorological conditions on the occurence of Cochlodinium polykrikoides and Pyrodinium bahamense var. compressum in coastal waters of Kota Kinabalu, Sabah, Malaysia. Sains Malaysiana, 43(1): 21–29 Montagnes D J S, Berges J A, Harrison P J, et al. 1994. Estimating carbon, nitrogen, protein, and chlorophyll a from volume in marine phytoplankton. Limnology and Oceanography, 39(5): 1044–1060. doi: 10.4319/lo.1994.39.5.1044 Omura T, Iwataki M, Borja V M, et al. 2012. Marine Phytoplankton of the Western Pacific. Tokyo: Kouseisha Kouseikaku Penna A, Casabianca S, Guerra A F, et al. 2017. Analysis of phytoplankton assemblage structure in the Mediterranean Sea based on high-throughput sequencing of partial 18S rRNA sequences. Marine Genomics, 36: 49–55. doi: 10.1016/j.margen.2017.06.001 Potvin M, Lovejoy C. 2009. PCR-based diversity estimates of artificial and environmental 18S rRNA gene libraries. Journal of Eukaryotic Microbiology, 56(2): 174–181. doi: 10.1111/j.1550-7408.2008.00386.x Ptacnik R, Solimini A G, Andersen T, et al. 2008. Diversity predicts stability and resource use efficiency in natural phytoplankton communities. Proceedings of the National Academy of Sciences of the United States of America, 105(13): 5134–5138. doi: 10.1073/pnas.0708328105 Řezanka T, Nedbalová L, Barcytė D, et al. 2019. Arsenolipids in the green alga Coccomyxa (Trebouxiophyceae, Chlorophyta). Phytochemistry, 164: 243–251. doi: 10.1016/j.phytochem.2019.05.002 Rodríguez F, Feist S W, Guillou L, et al. 2008. Phylogenetic and morphological characterisation of the green algae infesting blue mussel Mytilus edulis in the north and south Atlantic Oceans. Diseases of Aquatic Organisms, 81(3): 231–240 Rognes T, Flouri T, Nichols B, et al. 2016. VSEARCH: a versatile open source tool for metagenomics. PeerJ, 4: e2584. doi: 10.7717/peerj.2584 Roy R N. 1977. Red tide and outbreak of paralytic shellfish poisoning in Sabah. The Medical Journal of Malaysia, 31(3): 247–251 Sakamoto S, Lim W A, Lu Douding, et al. 2021. Harmful algal blooms and associated fisheries damage in east Asia: current status and trends in China, Japan, Korea and Russia. Harmful Algae, 102: 101787. doi: 10.1016/j.hal.2020.101787 Savin M C, Martin J L, LeGresley M, et al. 2004. Plankton diversity in the bay of fundy as measured by morphological and molecular methods. Microbial Ecology, 48(1): 51–65. doi: 10.1007/s00248-003-1033-8 Sidik M J, Rashed-Un-Nabi M, Hoque A M. 2008. Distribution of phytoplankton community in relation to environmental parameters in cage culture area of Sepanggar Bay, Sabah, Malaysia. Estuarine, Coastal and Shelf Science, 80(2): 251–260 Sildever S, Kawakami Y, Kanno N, et al. 2019. Toxic HAB species from the Sea of Okhotsk detected by a metagenetic approach, seasonality and environmental drivers. Harmful Algae, 87: 101631. doi: 10.1016/j.hal.2019.101631 Šlapeta J, López-García P, Moreira D. 2006. Global dispersal and ancient cryptic species in the smallest marine eukaryotes. Molecular Biology and Evolution, 23(1): 23–29. doi: 10.1093/molbev/msj001 Steidinger K A, Landsberg J H, Truby E W, et al. 1998. First report of Gymnodinium pulchellum (Dinophyceae) in north America and associated fish kills in the Indian River, Florida. Journal of Phycology, 34(3): 431–437. doi: 10.1046/j.1529-8817.1998.340431.x Stoeck T, Bass D, Nebel M, et al. 2010. Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Molecular Ecology, 19(S1): 21–31 Stoeck T, Hayward B, Taylor G T, et al. 2006. A multiple PCR-primer approach to access the microeukaryotic diversity in environmental samples. Protist, 157(1): 31–43. doi: 10.1016/j.protis.2005.10.004 Syasina I G, Kukhlevsky A D, Kovaleva A L, et al. 2012. Phylogenetic and morphological characterization of the green alga infesting the horse mussel Modiolus modiolus from Vityaz Bay (Peter the Great Bay, Sea of Japan). Journal of Invertebrate Pathology, 111(2): 175–181. doi: 10.1016/j.jip.2012.08.001 Teng S T, Abdullah N, Hanifah A H, et al. 2021. Toxic bloom of Pseudo-nitzschia cuspidata (Bacillariophyceae) and domoic acid contamination of bivalve molluscs in Malaysia Borneo. Toxicon, 202: 132–141. doi: 10.1016/j.toxicon.2021.09.018 Tillmann U, Gottschling M, Nézan E, et al. 2014. Morphological and molecular characterization of three new Azadinium species (Amphidomataceae, Dinophyceae) from the Irminger Sea. Protist, 165(4): 417–444. doi: 10.1016/j.protis.2014.04.004 Tomas C R. 1997. Identifying Marine Phytoplankton. San Diego: Academic Press Tyrrell J V, Bergquist P R, Gray R D, et al. 1996. Phylogeny of the raphidophytes Heterosigma carterae and Chattonella antiqua using ‘V4’ domain SSU rDNA sequences. Biochemical Systematics and Ecology, 24(3): 221–235. doi: 10.1016/0305-1978(96)00025-7 Vallina S M, Cermeno P, Dutkiewicz S, et al. 2017. Phytoplankton functional diversity increases ecosystem productivity and stability. Ecological Modelling, 361: 184–196. doi: 10.1016/j.ecolmodel.2017.06.020 Verma A, Hoppenrath M, Dorantes-Aranda J J, et al. 2016. Molecular and phylogenetic characterization of Ostreopsis (Dinophyceae) and the description of a new species, Ostreopsis rhodesae sp. nov., from a subtropical Australian lagoon. Harmful Algae, 60: 116–130. doi: 10.1016/j.hal.2016.11.004 Wilkerson F P, Grunseich G. 1990. Formation of blooms by the symbiotic ciliate Mesodinium rubrum: the significance of nitrogen uptake. Journal of Plankton Research, 12(5): 973–989. doi: 10.1093/plankt/12.5.973 Xu Xin, Yu Zhiming, Cheng Fangjin, et al. 2017. Molecular diversity and ecological characteristics of the eukaryotic phytoplankton community in the coastal waters of the Bohai Sea, China. Harmful Algae, 61: 13–22. doi: 10.1016/j.hal.2016.11.005 Yñiguez A T, Lim P T, Leaw C P, et al. 2021. Over 30 years of HABs in the Philippines and Malaysia: what have we learned?. Harmful Algae, 102: 101776 Yoshida K, Endo H, Lawrenz E, et al. 2018. Community composition and photophysiology of phytoplankton assemblages in coastal Oyashio waters of the western north Pacific during early spring. Estuarine, Coastal and Shelf Science, 212: 80–94 Yuan Jian, Li Meizhen, Lin Senjie. 2015. An improved DNA extraction method for efficient and quantitative recovery of phytoplankton diversity in natural assemblages. PLoS ONE, 10(7): e0133060. doi: 10.1371/journal.pone.0133060 Zamor R M, Glenn K L, Hambright K D. 2012. Incorporating molecular tools into routine HAB monitoring programs: Using qPCR to track invasive Prymnesium. Harmful Algae, 15: 1–7. doi: 10.1016/j.hal.2011.10.028 Zhu F, Massana R, Not F, et al. 2005. Mapping of picoeucaryotes in marine ecosystems with quantitative PCR of the 18S rRNA gene. FEMS Microbiology Ecology, 52(1): 79–92. doi: 10.1016/j.femsec.2004.10.006 -




 下载:
下载: